📋 Documentation Version Notice
You're reading documentation for v1.4, which is an older version. For up-to-date information, please have a look at v1.6 (Latest).
Analyzer → Importer Integration
Note
This integration is available from BIOMERO v2.3.0 / NL-BIOMERO v1.3.0 onward and requires
IMPORTER_ENABLED=true in your deployment.
Overview
Before this integration, analysis results (segmentation masks, measurements, etc.) produced by SLURM
workflows were imported back into OMERO exclusively via the OMERO API. This means the files landed on
OMERO server storage (the /OMERO volume), which grows without limit and bypasses the managed
remote-storage layer.
With the Analyzer → Importer integration the result files are instead:
Written to the shared remote storage that BIOMERO.importer already monitors.
Imported in-place by BIOMERO.importer — the same way manual data imports work — so they end up in the importer’s storage, not on the OMERO server volume.
Permanently archived in a dedicated
.analyzed/<workflow-uuid>/<timestamp>/folder, alongside the original zip, metadata CSV, and SLURM log file, for full traceability.
This makes analysis results a first-class citizen of the same storage and permission model as incoming research data, and avoids a split between “imported data” and “analysis results”.
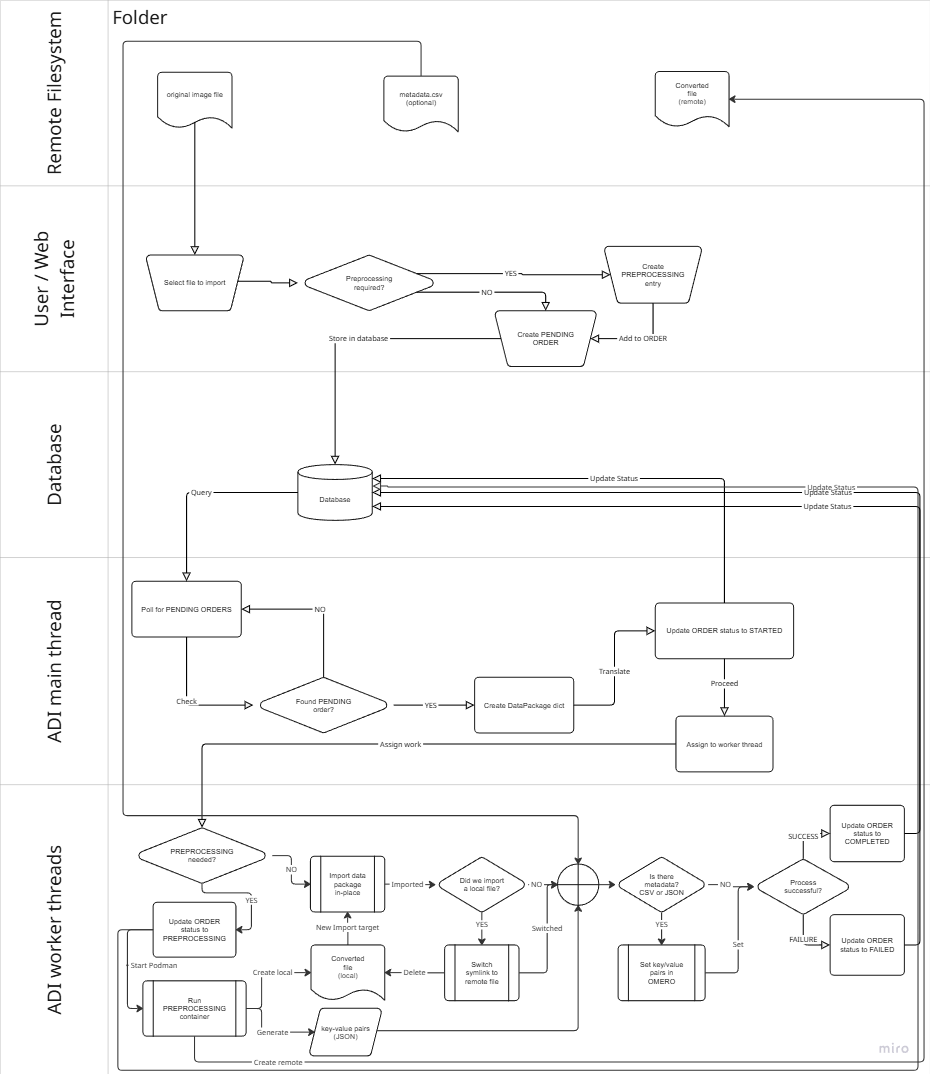
Analysis results are moved to remote (importer) storage and then imported in-place by BIOMERO.importer, using the same pipeline as manual imports.
How it works
Script selection
The BIOMERO worker selects the import script based on the IMPORTER_ENABLED environment variable:
IMPORTER_ENABLED=true→ runsSLURM_Import_Results.py(importer-enabled path)IMPORTER_ENABLED=false(default) → runsSLURM_Get_Results.py(classic OMERO API path)
Both scripts are bundled with BIOMERO.scripts and are present on the OMERO server container. The biomeroworker reads the env-var at runtime and passes the correct script name to the workflow orchestrator.
Permanent storage layout
Results are written to the shared drive under the group’s subfolder:
<group_base_path>/
└── .analyzed/
└── <workflow-uuid>/
└── <YYYYMMDD_HHMMSS>/
├── <job_id>_out.zip # original results archive
├── <job_id>_out/ # extracted result images + CSVs
│ ├── data/out/
│ │ ├── result_image.tiff
│ │ └── measurements.csv
│ └── metadata.csv # workflow provenance metadata
└── omero-<job_id>.log # SLURM job log
The .analyzed hidden directory distinguishes archived analysis results from raw incoming data.
Each workflow run gets its own <workflow-uuid>/<timestamp> directory, making multiple imports from
the same workflow safe.
Group base path resolution
The <group_base_path> is resolved in order:
Explicit group mapping in
biomero-config.json(same mappings used by the OMERO.biomero UI). If the user’s OMERO group has a configured folder mapping, that folder is used.Fallback:
<base_dir>/<group_name>wherebase_dircomes from the importer’ssettings.yml. For example, groupsystemwith no explicit mapping →/data/system/.
The group is taken from the OMERO user’s active group at the time they launched the analysis workflow. This means results always land in the group’s designated storage area.
Permission requirements
The BIOMERO worker process (running inside the biomeroworker container) must have write
permission on the group folder in the mounted storage:
If explicit mappings are configured, write permission is required on the mapped path.
If no mapping exists, write permission is required to create the group-named subfolder under
base_dir. For example, ifbase_dir=/dataand a user in groupteam-aruns their first workflow, the worker needs to be able to create/data/team-a/.
The default setup in NL-BIOMERO runs the biomeroworker as the omero-server user. Ensure this user
(or its UID, typically 999) has write access to the relevant directories on the host/NAS.
Upload order and in-place import
After copying result files to permanent storage, SLURM_Import_Results.py:
Creates an upload order in the BIOMERO.importer tracking database (
INGEST_TRACKING_DB_URL), pointing to the image files in.analyzed/….Polls the database until the order reaches
STAGE_IMPORTEDorSTAGE_INGEST_FAILED.Adds workflow metadata (key-value map annotations) to the imported images in OMERO once the import is confirmed.
BIOMERO.importer picks up the order, imports the files in-place, and updates the stage in the tracking database. The biomeroworker sees the status change via polling and reports back to the user.
Progress tracking
When track_workflows=True in slurm-config.ini, task status updates are visible in the
OMERO.biomero web interface:
IMPORTING— worker has written files to remote storage and submitted the upload order.IMPORTED— all upload orders confirmed as successfully imported by BIOMERO.importer.IMPORT_FAILED— one or more upload orders failed; check biomero-importer logs.
Infrastructure requirements
Environment variables for biomeroworker
The following env-vars must be set on the biomeroworker (in addition to the standard BIOMERO ones):
Variable |
Purpose |
Required |
|---|---|---|
|
Switches the import script to |
Yes |
|
PostgreSQL URL for the importer tracking database (same DB as BIOMERO.importer) |
Yes |
|
PostgreSQL URL for BIOMERO event-sourcing (workflow tracking) |
Yes (existing) |
Volume mounts for biomeroworker
Two additional volume mounts are required so the worker can read the importer configuration and the group-mapping JSON:
volumes:
# ... existing mounts ...
- "./config/biomero-importer:/opt/omero/server/config-importer:ro"
- "./web/biomero-config.json:/opt/omero/server/biomero-config.json:ro"
config/biomero-importer— containssettings.ymlfor BIOMERO.importer (same directory mounted into thebiomero-importercontainer). The worker readsbase_dirfrom this file.web/biomero-config.json— contains group-folder mappings used by OMERO.biomero. The worker uses these to resolve the correct destination path for each group.
Docker Compose changes
The biomeroworker service in docker-compose.yml now looks like:
biomeroworker:
build:
context: ./
dockerfile: ./biomeroworker/Dockerfile
args:
BIOMERO_VERSION: ${BIOMERO_VERSION}
BIOMERO_IMPORTER_VERSION: ${BIOMERO_IMPORTER_VERSION} # new
environment:
# ... existing ...
INGEST_TRACKING_DB_URL: postgresql+psycopg2://${BIOMERO_POSTGRES_USER}:${BIOMERO_POSTGRES_PASSWORD}@database-biomero:5432/${BIOMERO_POSTGRES_DB} # new
IMPORTER_ENABLED: ${IMPORTER_ENABLED}
volumes:
# ... existing ...
- "./config/biomero-importer:/opt/omero/server/config-importer:ro" # new
- "./web/biomero-config.json:/opt/omero/server/biomero-config.json:ro" # new
The Dockerfile itself installs biomero-importer (Python library, not the service) as a separate
layer so that it is available to the worker’s script execution environment:
RUN $VIRTUAL_ENV/bin/python -m pip install --ignore-requires-python \
biomero-importer==${BIOMERO_IMPORTER_VERSION}
Non-Docker deployments
If you are not using the NL-BIOMERO Docker Compose stack, you need to replicate the same setup manually on the BIOMERO worker host:
Install biomero-importer Python library in the same virtual environment as
biomero:pip install biomero-importer==<version>
Set environment variables before starting the OMERO processor:
export IMPORTER_ENABLED=true export INGEST_TRACKING_DB_URL=postgresql+psycopg2://user:pass@db-host:5432/biomero
Mount / configure file-system access so the worker process can write to the same storage location that BIOMERO.importer monitors.
Place configuration files at the paths the script expects:
settings.yml→/opt/omero/server/config-importer/settings.yml(or setIMPORTER_CONFIG_PATHif you customise this)biomero-config.json→/opt/omero/server/biomero-config.json
Permissions: ensure the worker process UID has write access to the group folders as described in Permission requirements above.
Troubleshooting
Results end up on OMERO server storage instead of remote storage
Check that
IMPORTER_ENABLED=trueis set on the biomeroworker container/process (not just on omeroweb or biomero-importer).Verify the biomero-importer Python library is installed in the worker’s virtual environment:
docker-compose exec biomeroworker \ /opt/omero/server/venv3/bin/python -c "import biomero_importer; print('ok')"
Upload order created but import never completes
Confirm biomero-importer container is running and healthy.
Check that the
INGEST_TRACKING_DB_URLon both the biomeroworker and the biomero-importer service point to the same database.Check biomero-importer logs for errors:
docker-compose logs biomero-importer
Files not found at expected path
Confirm all three containers (biomeroworker, biomero-importer, omeroserver) mount the storage at the same path (
/databy default); a mismatch will cause “file not found” errors at import time.Check that the group-folder mapping in
biomero-config.jsonpoints to a path that exists under the mounted storage root.
PermissionError writing to .analyzed folder
Verify the worker process UID has write access to the group base path on the host storage.
For new groups (first workflow run), the worker also needs permission to create the group subfolder under
base_dirif it does not exist yet.
Import polling times out
The default timeout is 3600 seconds (1 hour). For very large datasets the importer may take longer.
This timeout can be adjusted; please open an issue or customise
SLURM_Import_Results.pyfor your site.